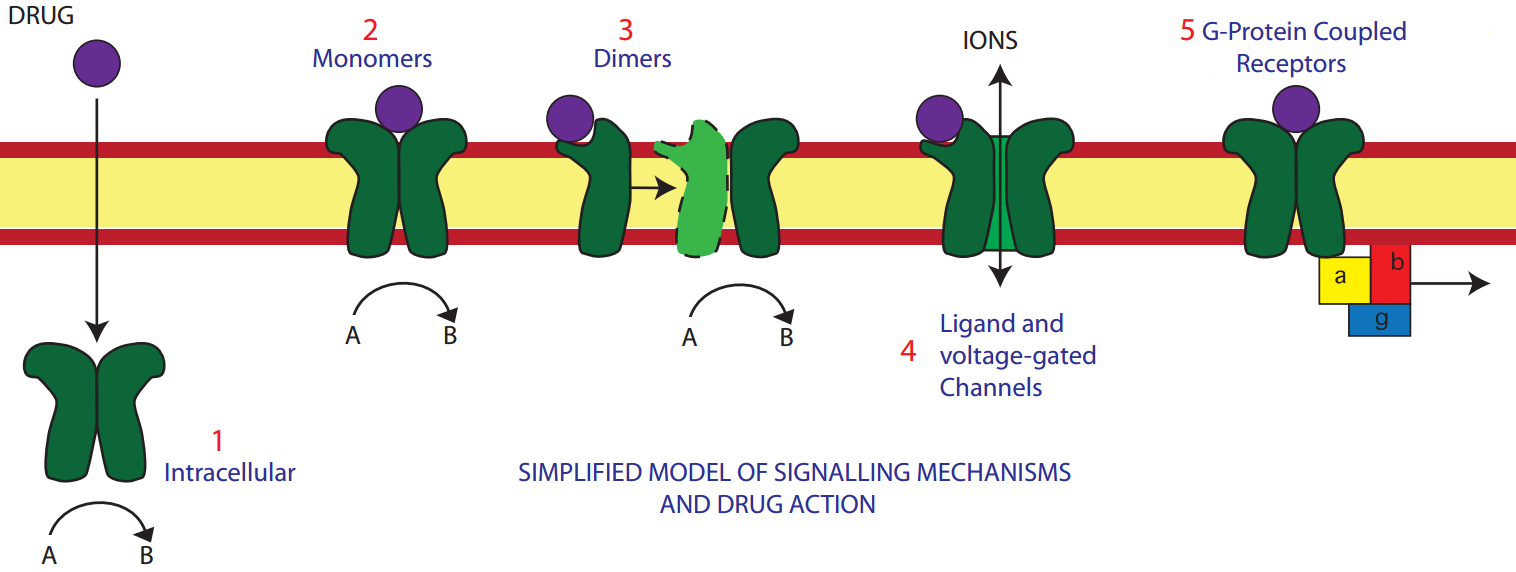
Receptor Types
To explain the concept of drug action with respect to: receptor theory, enzyme interactions, and physicochemical interactions.
To explain receptor activity with regard to: ionic fluxes, second messengers and G proteins, nucleic acid synthesis, evidence for the presence of receptors, regulation of receptor number and activity, structural relationships.
Receptor Types
A receptor is a protein, usually in the cellular membrane, to which a ligand may bind to generate a response.
- Intracellular receptors
May be either cytoplasmic or intra-nuclear.
- Enzyme-linked receptors
Are activated by a ligand and cause enzymatic activity on the intracellular side. They can be either:- Monomers
- Dimers
Where two proteins join, or diamerise, on binding of a ligand.
- Ion-channel receptors (ionotropic)
Create a channel through the membrane that allows electrolytes to flow down their electrical and concentration gradients. They can be either:- Ligand-gated channels
Undergo conformational change when a ligand is bound. There are three important families of ligand channels:- Pentameric family
Consist of five membrane spanning subunits. Include:- Nicotinic ACh receptor
- GABAA receptor
- 5-HT3 receptor
- Inotropic glutamate receptors
Bind glutamate, a CNS excitatory neurotransmitter. Include:
- NMDA receptor
High Ca2+ permeability
- NMDA receptor
- Inotropic purinergic receptors
Form cationic channels that are permeable to Ca2+, Na+, and K+
Activated by ATP
- Pentameric family
- Voltage-gated channels
Open when the threshold voltage is reached, facilitating electrical conduction in excitable tissues.- In their normal physiological state, voltage gated channels do not generally behave as receptors for a ligand, however some drugs (e.g. local anaesthetics) will bind to voltage gated channels to exert their effect
- Have a common 4-subunit structure (each with 6 transmembrane segments) surrounding a central pore
This pore is selective for the particular ion, which include:- Na+
- Located in myocytes and neurons
- Important in generating and transmitting an action potential by permitting sodium influx into cells
- Inhibited by local anaesthetics, anti-epileptics, and some anti-arrhythmics
- Ca2+
Divided into subtypes, including:- L
Muscular contraction. - T
Cardiac pacemaker. - N/P/Q
Neurotransmitter release. - K+
Located in myocytes and important in repolarisation following an action potential.
- L
- Na+
- Undergo a conformational change when the threshold potential is reached
This is sensed by the S4 helix, which acts to open and close the channel. - Exist in one of three functional states:
- Resting
Pore is closed. - Active
Pore is open, and ions can pass. - Inactive
Transient refractory period where the pore is open, but ions cannot pass. This creates the absolute refractory period of a cell.
- Resting
- Ligand-gated channels
- G-protein coupled (metabotropic) receptors:
G-proteins are a group of heterotrimeric (containing three units; α, β, γ) proteins which bind GDP. When stimulated, the GDP is replaced by GTP and the α-GTP subunit dissociates to activate or inhibit an effector protein. The effect depends on the type of α-subunit:- Gs proteins
Are stimulatorly. These- Increase cAMP, leading to a biochemical effect
- Gi proteins
Are inhibitory. These:- Inhibit adenylyl cyclase, reducing cAMP
- Gq proteins
Have a variable effect, depending on the cell. These:- Activate phospholipase C
This affects the production of:- Inositol triphosphate (IP3)
Stimulates Ca2+ from the SR, affecting enzymatic function or causing membrane depolarisation. - Diacylglycerol (DAG)
Activates protein kinase C, which has cell-specific effects.
- Inositol triphosphate (IP3)
- Activate phospholipase C
- Gs proteins
- Activate intracellular second messenger proteins when stimulated
Second messenger systems:- Result in both transmission and amplification of a stimulus, as a single activated receptor can activate multiple proteins and each activated protein may activate several other intermediate proteins
- This is known as a G-protein cascade
- Result in both transmission and amplification of a stimulus, as a single activated receptor can activate multiple proteins and each activated protein may activate several other intermediate proteins
Enzyme interaction
Drugs can interact with enzymes by antagonism or by being a false substrate.
Enzyme antagonism
Most drugs which interact with enzymes inhibit their activity. This results in:
- Increased concentration of enzymatic substrate
- Decreased concentration of the product of the reaction
Drugs can be competitive, non-competitive, or irreversible inhibitors of enzymatic activity. Examples include:
- Ramipril is a competitive inhibitor of angiotensin-converting enzyme.
- Aspirin is an irreversible inhibitor of cyclo-oxygenase.
False substrates
False substrates compete with the enzymatic binding site, and produce a product. Examples include:
- Methyldopa is a false substrate of the enzyme dopamine decarboxylase.
Physicochemical
Drugs whose mechanism of action is due to their physicochemical properties. Examples include:
- Mannitol reduces ICP because it increases tonicity of the extracellular compartment (and is unable to cross the BBB), drawing free water from the intracellular compartment as a consequence.
- Aluminium hydroxide reacts with stomach acid to form aluminium chloride and water, reducing stomach pH.
References
- Anderson, C. Pharmacodynamics 2. ICU Primary Prep.
- Law of Mass Action. Encyclopaedia Britannica.
- ANZCA August/September 2001
- Catterall WA. Structure and Function of Voltage-Gated Ion Channels. Annu. Rev. Biochem. 1995. 64:493-531.